Ashwini Rangaraj was one of the top 10 finalists at the Midwest 3MT competition! Congratulations on this massive achievement!
|
Sabrena gave an excellent oral presentation at the Society for Reproductive Investigation meeting in Vancouver in March. She also got the People's choice award for her talk at the Interdepartmental Biological Sciences Symposium! Congratulations, Sabrena!
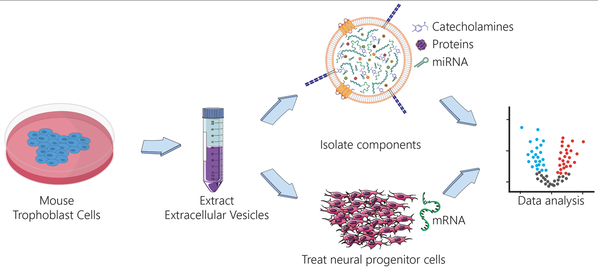
Check out or new collaborative manuscript where we looked at small RNA content of extracellular vessicles from mouse trophoblast stem cells (TSC) and TSC differentiated into parietal trophoblast giant cells (pTGC). We also looked at how exposure of mouse neural progenitor cells (NPC) to EV from either TSC or pTGC affect their transcriptome profiles. This exciting work is a first step towards understanding how placenta-derived extracellular vessicles may provide direct support for fetal brain development and may be an integral part of the placenta-brain axis. This work was in collaboration with Dr. Rosenfeld and Dr. Roberts from the University of Missouri, Columbia. Also check out this article about Cheryl, her work, and this published work!
Check out our collaborative manuscript where a conditional rat model targeting the invasive trophoblast cell lineage was generated. This is an excellent tool to study genes regulating invasive trophoblast cells in the placenta! This work was carried out in Dr. Soares group at KUMC.
Congratulations Ashwini Rangaraj whose abstract was selected for an oral presentation at the Greenwald Symposium! Ashwini gave an excellent talk about enhancer regulation in trophoblast cells. Congratulations also to Sabrena Rutledge, who presented a poster on her work and won the poster award!
Core conserved transcriptional regulatory networks define the invasive trophoblast cell lineage7/31/2023 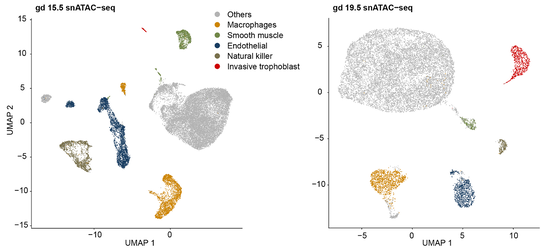 Our collaborative manuscript was published in Development today! We worked with the Michael Soares group to integrate our previously published single-cell RNA-seq data generated in the rat uterine-placental interface with single-nucleus ATAC-seq data generated at the same stage and in the same tissue type. We identified chromatin accessibility profiles in different cell types and identified a conserved gene regulatory network in invasive trophoblast cells. This work will will facilitate future studies investigating regulatory mechanisms in the invasive trophoblast cell lineage. 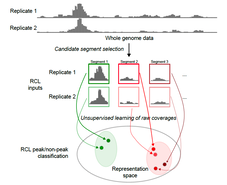 Our collaborative manuscript was published in Genome Research today! We worked with Yudi Zhang and Karin Dorman in the Statistics department on a novel peak caller that uses unsupervised contrastive learning to extract shared signals from multiple replicates. We focused on ATAC-seq data and demonstrated that our tool, replicative contrastive learner (RCL), outperformed other methods. Congrats all! |